Single-cell RNA-seq tool for neuronal cells Days 7 and 20 exposed to paracetamol
Here we present a searchable interactive web-tool utilising ShinyCell for the scRNA-seq analysis of neuronal differentiation time points Day 7 and 20 cells, control and after exposure to 100 and 200 micromolar paracetamol. This allows for exploration of gene expression in individual cells and annotated cell populations to explore the effect of drug exposure on gene expression.
A tool for single-cell ATAC-seq and RNA-seq integration data at Day 20 with and without paracetamol exposure
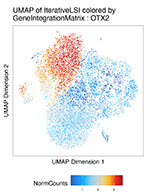
Explore chromatin opening in cells at neuronal cells at Day 20 that have been exposed to two theraputic doses of paracetamol during differentiation. The data has been analyzed using ArchR and is openly available in a webtool based on our in house ShinyArchR.UiO. Our webtool present different functionalities for genome-wide exploration of the single-cell datasets.
This data generated at the University of Oslo and is available in these tools under the permissive CC BY 4.0 license.
Citing our work: This work is published in iScience 10.1016/j.isci.2023.107755. The original preprint is available in biorxiv.org. The details of hESC differentiation is described in Star Protocols and our initial multiomics characterization of the neuronal differentiation protocol can be found in iScience. Code is available in EskelandLab Github. Processed datasets are deposited in NCBI GEO Superseries: GSE192858 and GSE220027.
