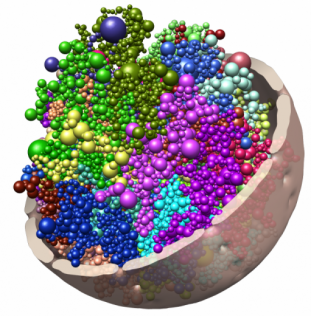
Chromatin is folded dynamically inside the nucleus, changing its conformation as cells divide and differentiate. The spheres in this image represent topologically associating domains (TADs), which can interact to form cliques that seem to stabilize heterochromatin at the nuclear periphery.
About the project
We are studying how topological domains and LADs, as genomic organizers, shape higher-order spatial genome topologies.
Ongoing research
- Computational methods for 3D and 4D modeling of genome architecture
- Functional relationship between 3D chromatin folding, radial positioning of chromatin, epigenetic states and lineage-specific differentiation
Outcomes / Recent findings
- Long-range associations between topological domains, forming cliques that shape the 4D genome during differentiation (Paulsen et al 2019 Nat. Genet 5, 835-843)
- Chrom3D genome structures in Virtual Reality (see our video on Youtube)
- UV-induced DNA lesions are enriched at the nuclear periphery (Garcia-Nieto et al 2017 EMBO J 36, 2829-2843)
- Chrom3D: a platform for 3D genome modeling from HiC and lamin-genome contacts (Paulsen et al 2018 Nat Prot 13, 1137-1152; Paulsen et al 2017 Genome Biol 18:21)
- ChIP protocol for nuclear lamins (Oldenburg et al 2016 Meth Mol Biol 1411, 315-324)
- Manifold-based optimization to enhance modeling of 3D chromatin structure from sparse HiC data (Paulsen et al 2015, PLoS Comput Biol 11, e1004396)
- Enriched Domain Detector (EDD): a domain calling to map LADs and other broad domains from ChIP-seq data (Lund et al 2014 Nucl Acids Res 42, e92)
Funding
The University of Oslo, The Research Council of Norway, The Norwegian Cancer Society.
Collaborations
- David Tremethick, The Australian National University, Canberra, Australia
- Lee Wong, Monash University, Clayton, Australia
- Steven Turner, Monash University, Clayton, Australia
- Ashby Morrison, Stanford University, Stanford, CA, USA